Research Overview
|
For my dissertation research in Ken Olsen's lab, I identified genetic and life history tradeoffs associated with local climatic adaptation in white clover (Trifolium repens). As part of this work, I assessed the relative importance of a well-studied chemical defense polymorphism (cyanogenesis), compared to other genomic factors, for climatic adaptation in this species.
I use field experiments, quantitative genetics, and population genomics approaches to characterize adaptive variation. My work addresses fundamental questions about the genetic architecture of local adaptation and the role of chemical defense polymorphisms in an obligately outcrossing perennial plant species. |
Being in white clover
|
|
Wikipedia: “To be (or to live) in clover” refers to living a carefree life of ease, comfort, and prosperity. This originally referred to the fact that clover is fattening to cattle.
As an evolutionary biologist, it has been a pleasure to study a species that grows (almost) everywhere! I frequently collect white clover seeds in my travels and have had great success using crowdsourcing to bolster my collections. The greenhouse staff at Washington University has been amazingly patient with my collections. They have helped me maintain, breed, and propagate thousands of plants over the past several years. I even built bee cages in the greenhouse, in which I released solitary bees to perform random pollination and generate F2 seeds! White clover has offered ample opportunities for mentoring undergraduate students. |
Population Genomics with Wild Samples
|
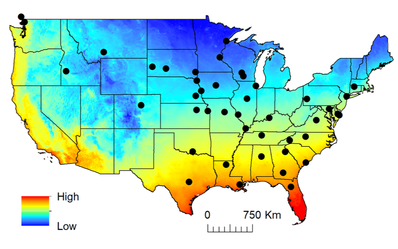
From my own travels and through seed crowdsourcing donations from numerous friends, family and colleagues, I have collected wild samples of white clover from 43 populations (416 total ecotypes, 7-10 per population) across North America.
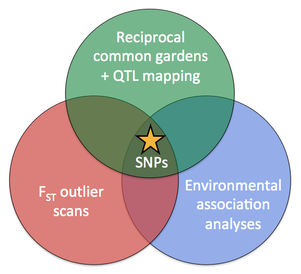
We are currently performing environmental association analyses (EAA) and FST outlier scans on genome-wide SNP markers to identify loci putatively associated with adaptation to local climates. |
Map colored by minimum winter temperature.
|
Field Experiments + QTL mapping
|
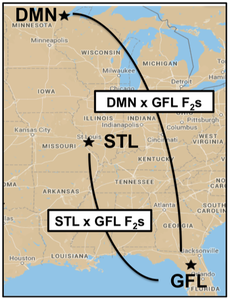
I planned and coordinated outdoor reciprocal common garden experiments at three field sites spanning a latitudinal gradient:
Duluth, MN (DMN), St. Louis, MO (STL), and Gainesville, FL (GFL). From 2016-2018, we measured fitness traits in the three gardens, which included 500 clonally-replicated F2 genotypes for two reciprocal comparisons. We used quantitative genetics and QTL mapping analyses to characterize the genetic architecture of local adaptation and to identify fitness-related genes/alleles in each environment. |
|
This research is funded by the National Science Foundation (NSF), including:
|
Rowan University · Department of Biological Sciences · 201 Mullica Hill Road · Glassboro, NJ 08028
wrights(at)rowan.edu · 856-256-4500 x53024
wrights(at)rowan.edu · 856-256-4500 x53024
Proudly powered by Weebly